| Version 69 (modified by , 12 years ago) ( diff ) |
|---|
Mussa
Multi-Species Sequence Analysis
"We be plundering the High Sequence Seas For the hidden Treasures of Conservation"
About Mussa
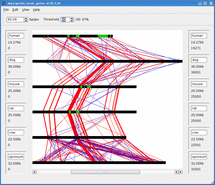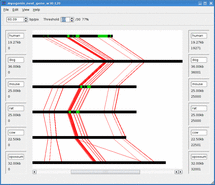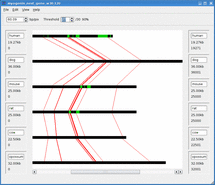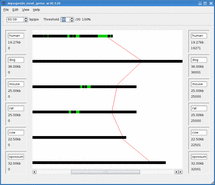
Mussa is a bioinformatics tool for finding and visualizing regions of conserved DNA by comparing across several species. It has been found that conserved regions are frequently enriched for regions that are functionally important. Thus Mussa is a tool to help discover previously unknown functional DNA elements.
Mussa's core algorithm is an N-way version of FamilyRelation's 2-way comparative sequence analysis software (which is a part of the Cartwheel project). Mussa analyzes the conservation between DNA sequence from N species by applying a transitivity requirement to all possible paths through all possible pairwise comparisons from the provided DNA segments to derive an N-way comparison.
For example, given sequences 1,2,3, and 4, Mussa makes 6 2-way comparisons: 1vs2, 1vs3, 1vs4, 2vs3, 2vs4, and 3vs4. It then compares all the links between these comparisons, saving those that satisfy a transitivity requirement. The saved paths are then displayed in an interactive viewer.
News
- 2009Nov23 - Mussa v1.1.0 (build 20091123 released)
- Updated to use Boost 1.40/1.41 and Qt 4.5.3.
- 2009Nov17 - Mussa Updates:
- We are in the process of updating Mussa to use Boost 1.40 and Qt 4.5.3.
- The Mac OS X and Windows binary builds are being tested and being prepared for release. The release will be Mussa 1.1.0.
- We will be switching the revision control system from bzr to git as well. Updated documentation for checkout of the source code will be included.
- 2007Jan24 - Release of the first Mussa Video Tutorial.
Download
Currently, we provide two types of binary releases for Mussagl. The first is a development build and the second is a stable release build.
Development Builds - New builds are made as new features / bug fixes are implemented. Many times these builds will be stable, but from time to time, a new feature will break something. Please report any bugs you find and hopefully we will be able to fix the bug and make a new build in a timely fashion.
Stable Releases - Stable releases are made when milestones have been reached and has been tested by our various test users.
Source - The source is available as well. Compiling from source is currently the only way to get a Linux build. Binary versions may be made available for Linux in the future, most likely Debian packages. See the bzr section of the wiki for information on downloading the latest source.
Stable Releases:
| Application | Version | Release Date | Mac OS X (Universal Binary) | Windows XP (Binary) | Source (Linux/Mac/Win) |
| Mussagl | 1.1.1 | Dec. 20 2013 | |||
| Mussagl | 1.1.0 | Nov. 23rd, 2009 | Download (.dmg) | Win32 Installer | Source (.tar.gz) |
| Mussagl | 1.0.0 | Nov. 1st, 2006 | Download (.dmg) | Win32 Installer | Source (.tar.gz) |
Contact
If you would like to contact us about Mussagl, feel free to e-mail one or both of us:
- Diane Trout <diane (at) caltech.edu>
- Brandon King <kingb (at) caltech.edu>
Documentation
- Mussagl User Guide
- FeatureNotes - documentation about specific features.
- Mussagl Video Tutorials
- Example Data
- Development Roadmap
- Checking out Mussagl from Revision Control
- Recent Mussagl Changes in Revision Control
- Building Mussagl from Source
Feature Requests / Bugs
Older Mussa Versions
- Mussa - C++/FLTK Version
- Mussa Prototype (Retired) - Python/PMW Version
Development Credits
- Mussagl: Diane Trout with assitance from Brandon King (Core algorithms retained from Mussa (C++/FLTK) )
- C++/FLTK conversion: Tristan De Buysscher with assistance from Titus Brown
- Python/PMW prototype: Tristan De Buysscher & Nora Mullaney
- Consulting User: Erich Schwarz
- Principal Investigators: Barbara Wold, Paul Sternberg
Attachments (1)
-
mussagl_animation.gif
(168.5 KB
) - added by 19 years ago.
Mussagl in action.
Download all attachments as: .zip